eGFRcrea GWAS and Prioritization
Kidney Epigenome and Transcriptome-based Multi-stage Prioritization for Kidney Disease
Welcome to eGFRcrea GWAS and Prioritization Atlas
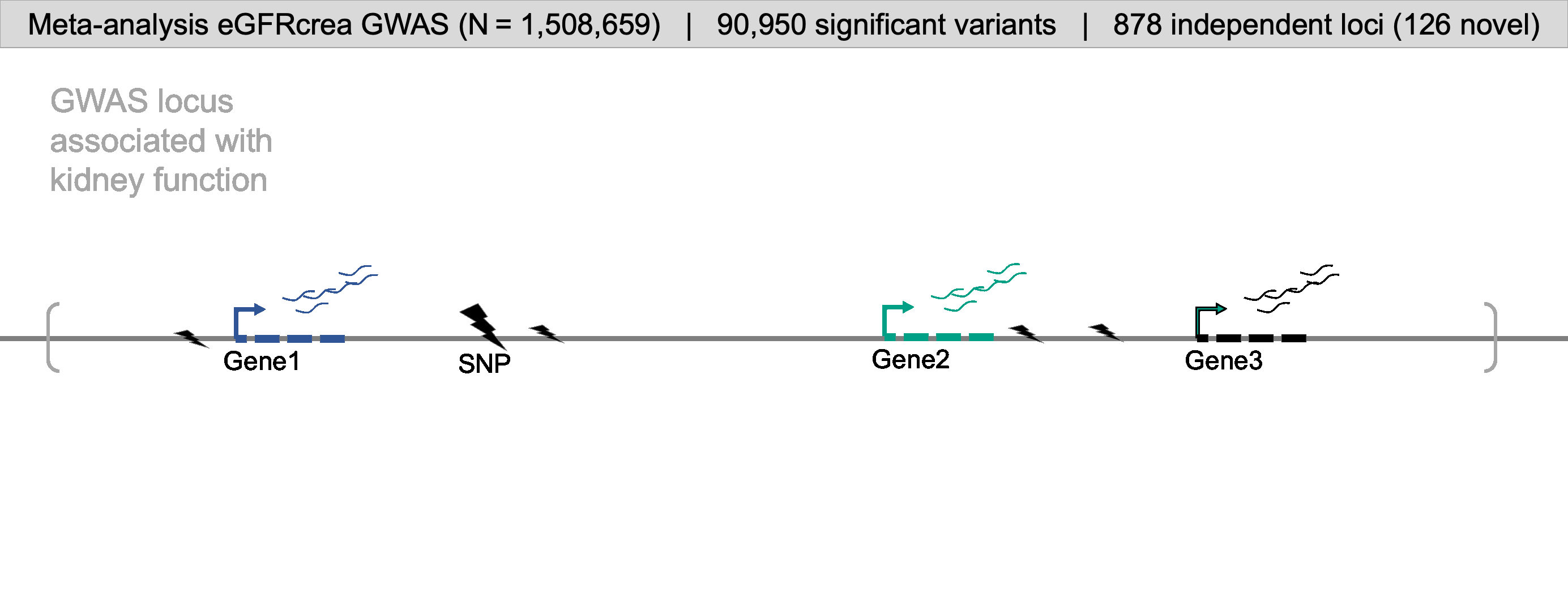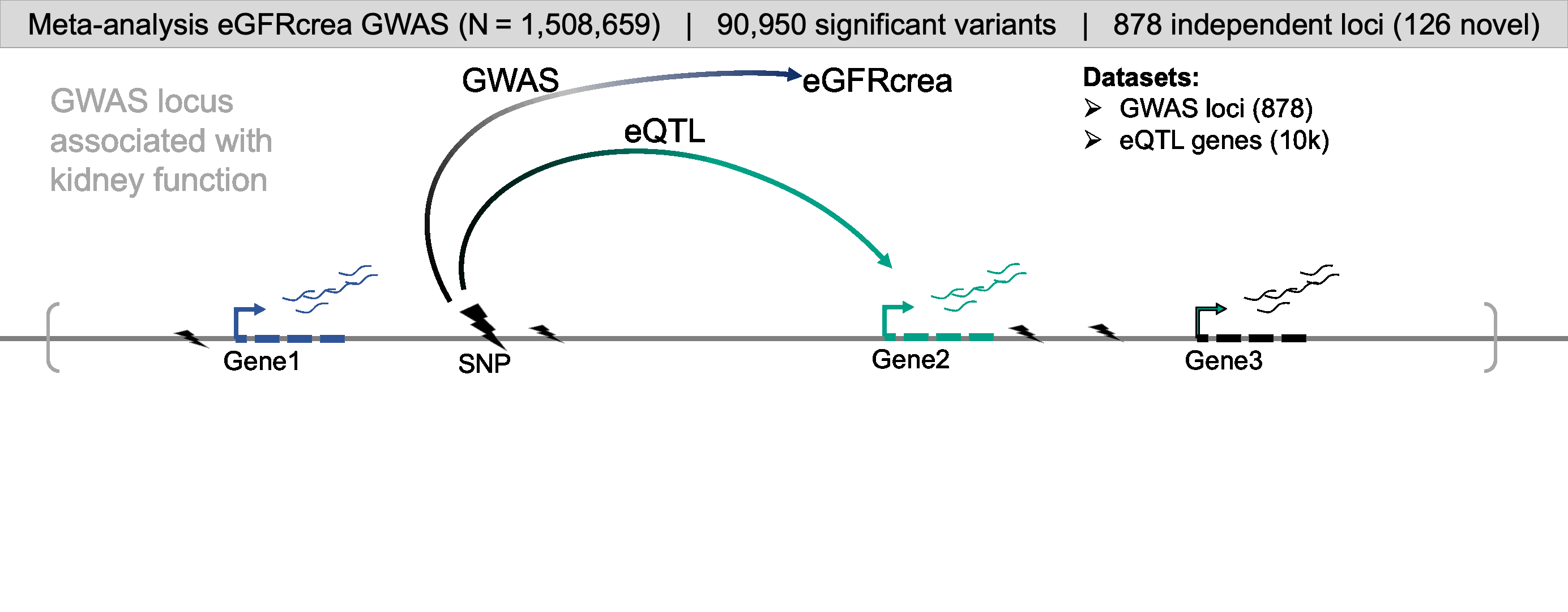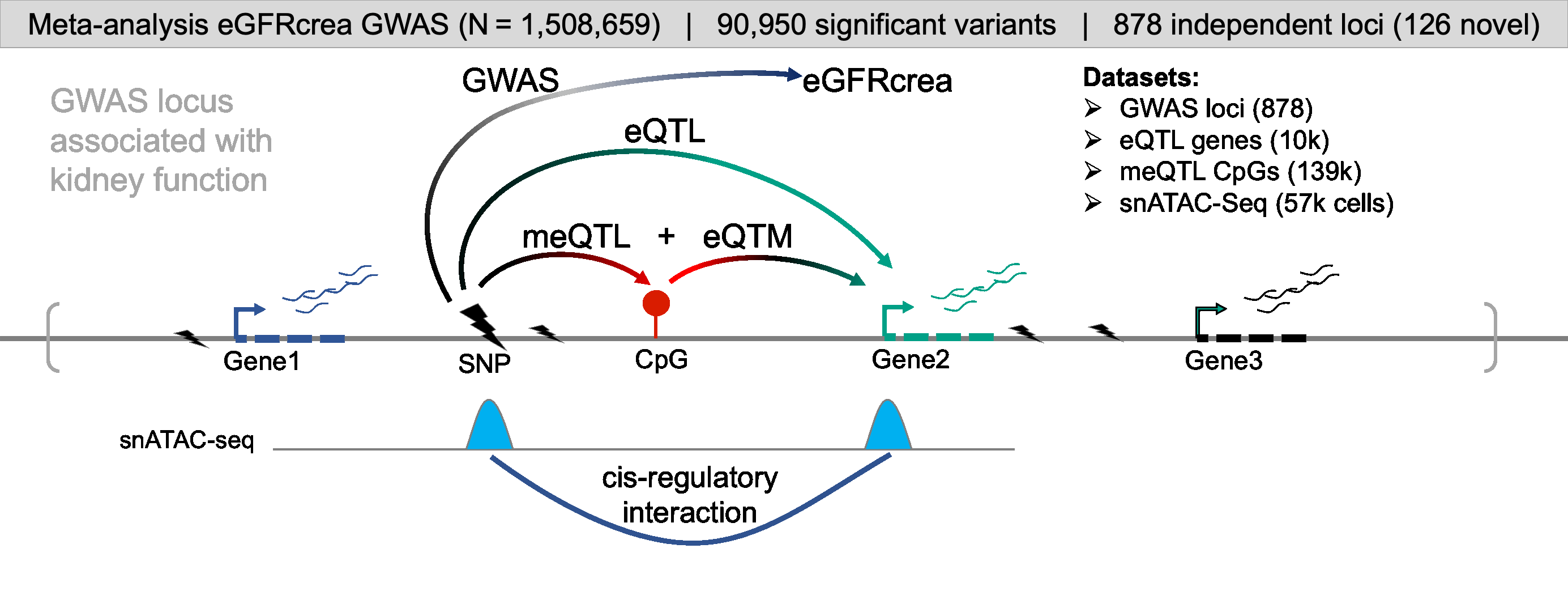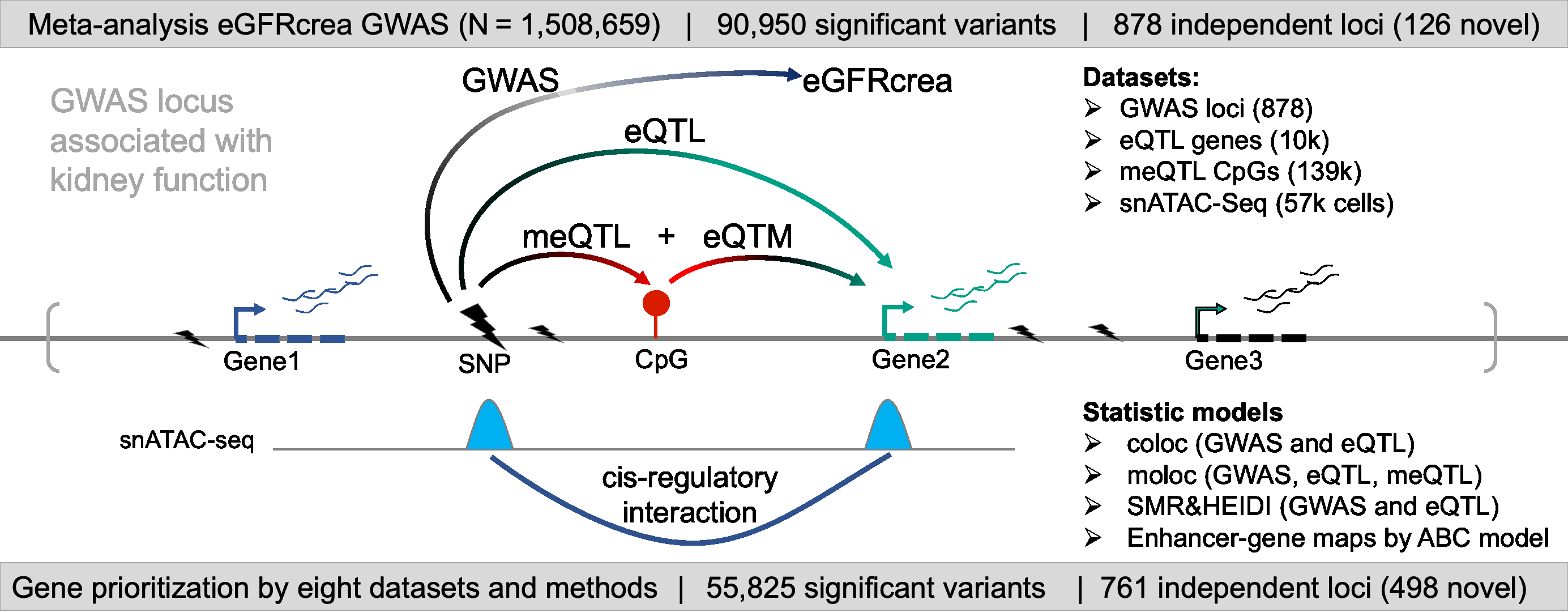
For comprehensive identification of genetic variants associated with kidney function (eGFRcrea: glomerular filtration rate estimated by serum creatinine), we conducted a meta-analysis of publicly available GWAS information obtained from CKDGen, Pan-UK Biobank, MVP, PAGE, and SUMMIT consortium. GWAS meta-analysis resulted in a comprehensive eGFRcrea GWAS catalog of 12 million variants with a total sample size of more than 1.5 million (GWAS Catalog). Totally, we identified 878 independent loci (including 126 novel loci), most of which were validated by eGFRcys and/or BUN. Further, we developed a prioritization scoring system to identify target genes for kidney function GWAS loci by integrating evidence from eight different omics datasets or analytical tools, such as eQTL, meQTL and eQTM, coloc (GWAS and eQTL), moloc (GWAS, eQTL and meQTL), SMR, HEIDI, Cicero connection, and enhancer-promoter contacts by Activity-by-Contact Model. This strategy enabled us to prioritize target genes for 761 (86.7% from total 878) independent loci.