Human Kidney eQTL Atlas
Mapping the genetic architecture of human traits to cell types in the kidney identifies mechanisms of disease and potential treatments
Welcome to the Susztaklab human kidney eQTL atlas
The functional interpretation of GWAS remains challenging due to the cell-type dependent influences of genetic variants. To comprehensively annotate the genotype effect on gene expression in the kidney, we conducted several eQTL analysis in human kidneys.
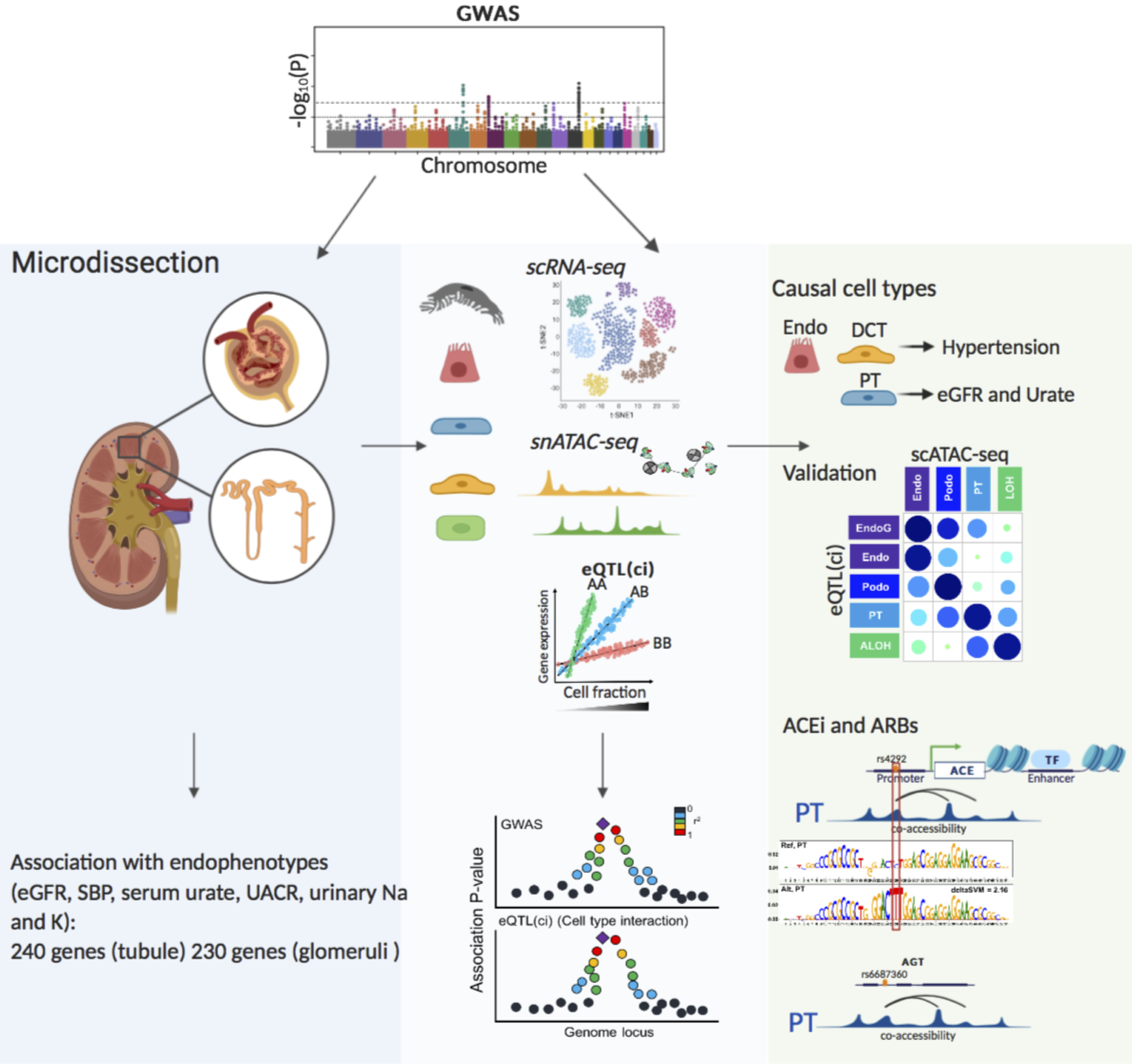
Sheng et al. generated comprehensive maps of expression quantitative trait loci (eQTL) for 659 microdissected human kidney samples and identified cell-type eQTLs by mapping interactions between cell type abundance and genotype. By partitioning heritability using stratified LD-score regression to integrate GWAS with scRNA-seq and snATAC-seq data, we prioritized proximal tubules in kidney function and endothelial cells and distal tubule segments in blood pressure pathogenesis. Bayesian colocalization analysis nominated more than 200 genes for kidney function and hypertension. Our study clarifies the mechanism of commonly used antihypertensive and renal protective drugs and identifies drug repurposing opportunities for kidney disease.
In our latest study, Liu et al. performed meta-analysis by integrating all publicly available kidney eQTL data (Sheng et al., Ko et al., GTEx v8, and NephQTL). The meta-analysis eQTL dataset included 201,627,059 associations with a total sample size of n = 686. Based on this new database, we identified 10,430 genes with eQTLs, of which 11% (1,146) have not been previously reported. This most comprehensive eQTL dataset enabled us prioritize 66% of GWAS loci associated with kidney function.
 Kidney eQTLs (eQTL(meta)) identified by meta-analysis of four studies with total 686 human kidney samples are available online.
Kidney eQTLs (eQTL(meta)) identified by meta-analysis of four studies with total 686 human kidney samples are available online.